The page illustrates how to run the PCMDI metrics package (PMP) on ACME model output. Please be aware that PMP is still experimental and under development. Currently, PMP is only supported on crunchy.llnl.gov.
Prerequisites
- Account on crunchy.llnl.gov
- Climatology files generated by ncclimo (Generating and Regridding Climatologies (climo files) with NCO and ncclimo). Sample files are available on crunchy for the step-by-step guide below.
Step-by-step guide
Load environment
|
|---|
Use cdscan to create xml describing input files
|
|---|
The output file be a file named xmls/alpha6_1850_0070-0099.xml that describes the input netcdf files.
Process climatologies to be consistent with CMIP data structure
|
|---|
The result will be a number of individual netcdf files in the sub-directory data/alpha6_1850_0070-0099:
ls data/alpha6_1850_0070-0099/
pr.nc prw.nc psl.nc rlutcs.nc rlut.nc rsutcs.nc rsut.nc ta.nc tas.nc tauu.nc tauv.nc ua.nc va.nc zg.nc
Compute derived fields (currently only CRE fields)
|
|---|
Two additional netcdf files for LW and SW cloud radiative effects will be created under data/alpha6_1850_0070-0099:
ls data/alpha6_1850_0070-0099/pr.nc prw.nc psl.nc rltcre.nc rlutcs.nc rlut.nc rstcre.nc rsutcs.nc rsut.nc ta.nc tas.nc tauu.nc tauv.nc ua.nc va.nc zg.nc
Run PMP mean climate statistics
|
|---|
This is the core of PMP. It will compute climate statistics comparing the model output with various observational datasets. The statistics are stored in json files located under the subdirectory results/pmp:
ls results/pmp/*.json
results/pmp/pr_2.5x2.5_esmf_linear_metrics.json results/pmp/ta-850_2.5x2.5_esmf_linear_metrics.json
results/pmp/prw_2.5x2.5_esmf_linear_metrics.json results/pmp/tas_2.5x2.5_esmf_linear_metrics.json
results/pmp/psl_2.5x2.5_esmf_linear_metrics.json results/pmp/ua-200_2.5x2.5_esmf_linear_metrics.json
results/pmp/rltcre_2.5x2.5_esmf_linear_metrics.json results/pmp/ua-850_2.5x2.5_esmf_linear_metrics.json
results/pmp/rlut_2.5x2.5_esmf_linear_metrics.json results/pmp/va-200_2.5x2.5_esmf_linear_metrics.json
results/pmp/rstcre_2.5x2.5_esmf_linear_metrics.json results/pmp/va-850_2.5x2.5_esmf_linear_metrics.json
results/pmp/ta-200_2.5x2.5_esmf_linear_metrics.json results/pmp/zg-500_2.5x2.5_esmf_linear_metrics.json
There is one json file for every variable. Additionally, re-gridded netcdf files are under the subdirectory interpolated-output.
Create portrait plots
|
|---|
The resulting output should be a png image file under the subdirectory plots:
ls plots
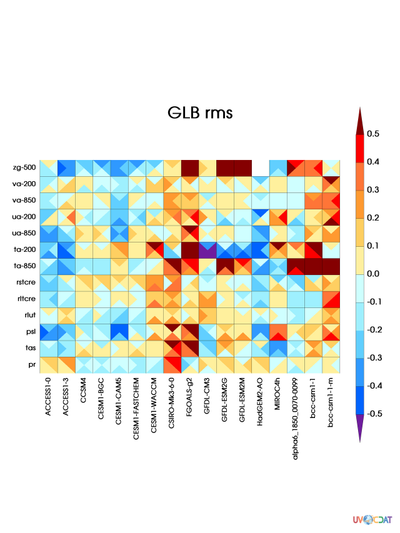
acme_pmp-control_tests_rms_GLB.png
This image should look like
To generate portrait plots for other regions (tropics, nothern extra-tropics, souther extra-tropics), edit portrait_plots.py and change definition of the region from global to another option and then rerun the script.
reg = 'GLB' #reg = 'NHEX' #reg = 'SHEX' #reg = 'TROPICS' |
|---|
Questions?
Chris Golaz may be able to help.
Acknowledgements
Many thanks to Peter Gleckler and Charles Doutriaux (Unlicensed) for their help getting me started with PMP.